[author: Kevin E. Noonan]
 Earlier this month we asked for examples of instances where claims to isolated DNA interfered with basic research (see "The Proper Scope of DNA (or "Gene") Patent Claims"). We posited that the claim would be something like this one:
Earlier this month we asked for examples of instances where claims to isolated DNA interfered with basic research (see "The Proper Scope of DNA (or "Gene") Patent Claims"). We posited that the claim would be something like this one:
An isolated (human) nucleic acid (or DNA molecule or gene (specified by name)), encoding an amino acid sequence identified by SEQ ID NO: X,
where the portions of this claim in parentheses are optional, used in some cases but not necessary to properly recite the claimed subject matter. This type of claim has the following features:
• It must use the term "isolated" in the claim, to distinguish it from naturally occurring molecules in their natural state, so as to evince the "hand of man" in its creation (Diamond v. Chakrabarty) to satisfy 35 U.S.C. § 101;
• It cannot read on the natural molecule (i.e., the claim scope cannot be broad enough to encompass the natural molecule) so that it satisfies the novelty requirement of 35 U.S.C. § 102 (Wood-Paper Patent, 90 U.S. 566 (1874), and Cochrane v. Badische Anilin Soda Fabrik, 111 U.S. 293 (1884));
• Infringement will lie only if the entire amino acid sequence recited in the claim is made, used, sold, offered to be sold, or imported;
• Portions of the gene can be isolated, sequenced, and characterized and not infringe, and such portions can be changed and then introduced into the gene to produce another gene that does not infringe;
• The gene product (typically, a protein) can be isolated from the cell and studied without infringement, as can antibodies raised against the gene product; and
• The sequence itself can be identified in individuals and compared to wildtype or disease-associated mutations without infringing unless the entire DNA is isolated intact.
 Recent posts provide an opportunity to further distinguish what is patent eligible from what is not (see "Dolphin Genes Show Relationships between Large Brains and Energy Metabolism Similar to Humans and Elephants" and "Genetic Marker Map of Sunflower Species Reported"). In the former, researchers have identified a number of genes that have been subjected to positive selection, presumably due to their usefulness in adapting to the aquatic lifestyle. This information was developed by comparing dolphin genes with their orthologs in related species, including cow, horse, dog, mouse, elephant, opossum, platypus, chicken, and man. And none of this information is, per se, eligible for patenting, being (like the capacity for certain species of Rhizobium to grow together) to be "part of the storehouse of knowledge of all men [and] manifestations of laws of nature, free to all men and reserved exclusively to none." Funk Brothers Seed Co. v. Kalo Inoculant Co., 333 U.S. 127 (1948).
Recent posts provide an opportunity to further distinguish what is patent eligible from what is not (see "Dolphin Genes Show Relationships between Large Brains and Energy Metabolism Similar to Humans and Elephants" and "Genetic Marker Map of Sunflower Species Reported"). In the former, researchers have identified a number of genes that have been subjected to positive selection, presumably due to their usefulness in adapting to the aquatic lifestyle. This information was developed by comparing dolphin genes with their orthologs in related species, including cow, horse, dog, mouse, elephant, opossum, platypus, chicken, and man. And none of this information is, per se, eligible for patenting, being (like the capacity for certain species of Rhizobium to grow together) to be "part of the storehouse of knowledge of all men [and] manifestations of laws of nature, free to all men and reserved exclusively to none." Funk Brothers Seed Co. v. Kalo Inoculant Co., 333 U.S. 127 (1948).
However, the paper does posit that dolphins may be large animal models for type 2 diabetes in humans
During the course of primate evolution, metabolism evolved to provide a constant supply of energy to the increasing demands of a larger brain. [Leonard et al., 2003, "Metabolic correlates of hominid brain evolution," Comp. Biochem. Physiol. A 136: 5–15]. Humans have evolved several adaptations, including an increase in visceral fat and the evolution of tissue-specific insulin resistance in the brain that protects nervous tissue from energy shortage; however, malfunctions of this system in humans can cause type 2 diabetes (Kuzawa, 2010 Beyond feast-famine: brain evolution, human life history, and the metabolic syndrome. In Human evolutionary biology (ed. Muehlenbein M.), pp. 518–527. Cambridge, UK: Cambridge University Press). Interestingly, recent research has proposed that dolphins should be seen as a model for type 2 diabetes, as they possess all the hallmarks of the disease [Venn-Watson et al., 2011, "Dolphins as animal models for type 2 diabetes: sustained, post-prandial hyperglycemia and hyperinsulinemia," Gen. Comp. Endocrinol. 170: 193–99)]. Here, we find genomic signatures of adaptive evolution in dolphin genes related to control of food intake such as genes involved in glycerol uptake and/or glucose metabolism (AQP9, OSTN, SOCS6), and a neuropeptide AGRP that is expressed in the hypothalamus and involved in increasing appetite, decreasing metabolism and regulating leptin. In addition, we found several genes under selection related to lipid transport and metabolism [] that may be associated with the large fat reserves found in cetaceans.
Thus, methods for testing compounds as potential drugs for treating human type 2 diabetes using dolphins may be patent eligible, in a claim such as this one:
A method for identifying a compound that is effective in treating type 2 diabetes in the human, comprising the step of administering the compound to a bottlenose dolphin and detecting an antiglycemic effect in the dolphin.
The other paper, on the sunflower genome identified more than 10,000 genetic loci (by virtue of small nucleotide polymorphisms or SNPs) to provide a genetic map of the sunflower genome. That genome was represented graphically as follows:
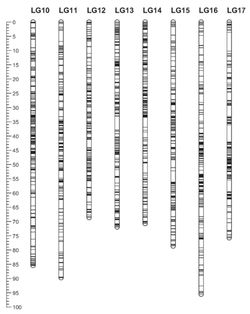


 Like the dolphin genome, this map is also not patent eligible. However, sunflowers are of great economic importance, being the fourth largest source of cultivated vegetable oils worldwide (www.fas.usda.gov). In addition to these maps, the "sunflower genome project" is under active pursuit (Kane et al. 2011, "Progress towards a reference genome for sunflower," Botany-Botanique 89: 429–37), wherein genes that may increase yield or resistance to insect predation or aridity may be mapped to portions of the sunflower genome close to one of these SNPs. In such a case, methods for identifying sunflowers having a desirable trait associated with one of these polymorphisms may be patentable:
Like the dolphin genome, this map is also not patent eligible. However, sunflowers are of great economic importance, being the fourth largest source of cultivated vegetable oils worldwide (www.fas.usda.gov). In addition to these maps, the "sunflower genome project" is under active pursuit (Kane et al. 2011, "Progress towards a reference genome for sunflower," Botany-Botanique 89: 429–37), wherein genes that may increase yield or resistance to insect predation or aridity may be mapped to portions of the sunflower genome close to one of these SNPs. In such a case, methods for identifying sunflowers having a desirable trait associated with one of these polymorphisms may be patentable:
A method for producing sunflowers having [a desirable trait], wherein the trait is genetically linked to [a particular SNP], the method comprising the step of identifying the SNP in said sunflower and cultivating said sunflower.
These examples are not meant to be exhaustive. But they are intended to illustrate the difference between patent eligible and ineligible subject matter, and to show that the mere intersection of patents and biology should not be the occasion to disqualify patent eligible subject matter.