The advent of technology making feasible elucidation of whole genomic sequencing over the past 30 years has led to reports of many if not most important or interesting animal genomes (including the most celebrated results of the Human Genome Project) (see, e.g., "Nautilus pompilius Genome Determined"; "Giraffe Genome Reveals Relevant Adaptations"; "Avocado Genome Elucidated; Durum Wheat Genome Revealed"; "Rose Genome Reveals Its Exquisite Complexities"; "Silver Birch Genetics Explained"; "Genomic Sequence of Strawberry Determined"; "Koala Genome Sequenced"; "Tomato Genome Determined"; "Lowland Gorilla Genome Sequenced"), as well as important historical relationships newly appreciated (see, e.g., "Did Neanderthal DNA Persist in Modern Humans as a Defense against Xenobiotic Viruses?"; "Chicken Origins Established (But Philosophical Questions Remain)"; "Genetic Analyses of Sweet Potato Genome Sheds Light on Speciation and Global Dispersion Patterns"; "The Domestication History of Apples Revealed by Genomic Analysis"; "Evidence of Geographic Change in Central America from Genome Studies of Eciton Ant Species"; "Dolphin Genes Show Relationships between Large Brains and Energy Metabolism Similar to Humans and Elephants").
 But what has become evident recently is that most of these genomes were in one way or another incomplete, due typically because there were regions of this or that genome that were resistant to accurate sequencing or come other biologic idiosyncrasy. The most recent example of this reminder of the complexity of biological organisms is our old friend the domestic cat, which built (as most of the new studies do) on the earlier genomic elucidations. In this case, the earlier study in question was published in 2014, when an international effort led by Stephen J. O'Brien at the Oceanographic Center, Nova Southeastern University, Ft. Lauderdale, Florida reported the complete genomic sequencing of the domestic cat, Felix catus. The report, entitled "Annotated features of domestic cat – Felis catus genome," was published in GigaScience 2014, 3:13 (August 5, 2014) (see "Domestic Cat Genome Sequenced"). The study reported sequencing of a female Abyssinian cat named Cinnamon, a mixed-breed cat from Russian named Boris, and Sylvester, a wildcat ancestor of domestic cats. The report showed that domestic cats have retained "a highly conserved ancestral mammal genome organization" in comparison with ancestral cats (see Driscoll et al., 2007, "The near eastern origin of cat domestication," Science 317: 519–23). Both species, F. catus and Felix silvestris, have 38 chromosomes, 18 pairs of autosomes, and two pairs of dimorphic gender-determining chromosomes. Details of the domestic cat genome structure included the presence of 217 loci of endogenous retrovirus-like elements (amounting to 55.7% of the entire genome, comprised of long interspersed elements (LINEs), short interspersed elements (SINEs), satellite DNA, retroviral long terminal repeats (LTRs) and "others"); 21,865 protein coding genes (open reading frames or ORFs), detected by comparison with eight mammalian genomes (from human, chimpanzee, macaque, dog, cow, horse, rat, and mouse); and a wealth of genetic variability in single nucleotide polymorphisms (SNPs), insertion/deletion events (indels); novel families of complex tandem repeat elements; and short terminal repeat (STR) loci.
But what has become evident recently is that most of these genomes were in one way or another incomplete, due typically because there were regions of this or that genome that were resistant to accurate sequencing or come other biologic idiosyncrasy. The most recent example of this reminder of the complexity of biological organisms is our old friend the domestic cat, which built (as most of the new studies do) on the earlier genomic elucidations. In this case, the earlier study in question was published in 2014, when an international effort led by Stephen J. O'Brien at the Oceanographic Center, Nova Southeastern University, Ft. Lauderdale, Florida reported the complete genomic sequencing of the domestic cat, Felix catus. The report, entitled "Annotated features of domestic cat – Felis catus genome," was published in GigaScience 2014, 3:13 (August 5, 2014) (see "Domestic Cat Genome Sequenced"). The study reported sequencing of a female Abyssinian cat named Cinnamon, a mixed-breed cat from Russian named Boris, and Sylvester, a wildcat ancestor of domestic cats. The report showed that domestic cats have retained "a highly conserved ancestral mammal genome organization" in comparison with ancestral cats (see Driscoll et al., 2007, "The near eastern origin of cat domestication," Science 317: 519–23). Both species, F. catus and Felix silvestris, have 38 chromosomes, 18 pairs of autosomes, and two pairs of dimorphic gender-determining chromosomes. Details of the domestic cat genome structure included the presence of 217 loci of endogenous retrovirus-like elements (amounting to 55.7% of the entire genome, comprised of long interspersed elements (LINEs), short interspersed elements (SINEs), satellite DNA, retroviral long terminal repeats (LTRs) and "others"); 21,865 protein coding genes (open reading frames or ORFs), detected by comparison with eight mammalian genomes (from human, chimpanzee, macaque, dog, cow, horse, rat, and mouse); and a wealth of genetic variability in single nucleotide polymorphisms (SNPs), insertion/deletion events (indels); novel families of complex tandem repeat elements; and short terminal repeat (STR) loci.
The report also contained an extensive comparison between domestic cats and other species ("reference genomes") in terms of gene numbers, using genes with the longest mRNA and corresponding coding sequences (click on table to expand).

More recently, further results of applications of improved genomic sequencing methods and technologies have provided a more comprehensive elucidation of the feline genome and insights into genetic bases for disease. A paper published in the Public Library of Science, entitled "A new domestic cat genome assembly based on long sequence reads empowers feline genomic medicine and identifies a novel gene for dwarfism," PLoS Genetics 16(10): e1008926 on October 20, 2020, reported a revisit of the genomic sequence of Cinnamon, an Abyssinian breed domestic cat previously sequenced. Rather than focusing on one cat, this group* performed whole genome sequencing (WGC) of 54 domestic cats and aligned the sequences to detect single nucleotide variants (SNVs) and structural variants (SVs). The distribution and relatedness of the cats in this study is shown in this graphic:

Their aim was to identify the sequence comprising the ~300,000 gaps in the annotated sequence reported to the Cinnamon Abyssinian, to produce a new reference cat genome denoted in relevant databases as Felis_catus_9.0. This genome comprised 2.84 gigabasepairs (Gb), of which only 1.8% (1.38 megabasepairs, Mb) was not assigned to a specific chromosomal location. The sequencing and comparison identified 19,748 genes, 376 of which were novel (i.e., had not been identified in Cinnamon's DNA) and 178 genes found in the original annotations but not found here. (The authors note the presence of 54 diverse individual cats from which the genes were determined as a possible source of this disparity.) A greater extent of LINE/L1 repetitive elements were detected, which these researchers attributed to better assembly (i.e., the repetitiveness and gaps did not mask these elements in the latest assembly). The average number of SNVs detected per cat was 9.6 million (for comparison humans have ~5 million); inbred cats showed a lower number (~8 million) while outbred cats showed a higher number (> 10.5 million). Of these, researchers detected 128,844 synonymous SNVs in protein-coding sequences (i.e., where the two sequence encoded proteins having the same amino acid sequence), 77,662 missense SNVs (wherein the protein encoded differed in amino acid sequence, including truncated sequences and out-of-frame mutants), and 1,179 loss-of-function (LoF) SNVs. LoF and missense SNVs were associated with depletion in comparisons between orthologous cat and human DNA sequences, whereas synonymous SNVs were enriched by 19.2%. Statistical analyses found that in cats and humans there was selection against missense and LoF SNVs in genes under selective pressure (i.e., where the genetic changes resulted in phenotypes deleterious to individuals carrying them). These results further indicated that these classes of SNVs were not distributed randomly in the cat or human genome.
Turning to structural variants, these researchers detected an average of 44,900 SVs per cat (a frequency four-fold higher than in humans), comprising 134.3 Mb; of the structural variant types, deletions averaged 905 bp, duplications 7,497 bp, insertions 30 bp, and inversions 10,993 bp. In total they reported 208,135 SVs detected in 54 sequenced cat genomes, with 123,731 (60%) were deletions, approximately 39,955 insertions, 35,427 inversions, and 9,022 duplications were identified. The majority of these were common across cat breeds, indicating tolerance. 58.15% of these SVs were intergenic in location, 40.22% intronic, and only 1.06% found in exons, "potentially impacting 217 different protein coding genes. As the scientists noted:
Conversely, the proportion of some SV types found in certain gene regions varied from their genome-wide averages. For example, in regions 5 kb upstream and downstream of genes, duplications were increased approximately two-fold. For exonic regions, 74% of SVs were deletions, an increase from the genome wide level of 59.45%. For 5' untranslated regions (UTRs), the majority of SVs were inversions, which only represent 17.02% of total SVs. These results suggest an interaction between the impact of SV types and the potential function of the gene regions they are found in.
Sixteen of the identified variants were predicted to cause disease, based on comparisons with human genome sequences known to be sufficiently similar. These included one involving a tumor suppressor gene FBXW7 associated with feline mediastinal lymphoma, prevalent in Siamese cats and Oriental shorthaired cats. Other variants detected and their related diseases were in the FAM13B gene, resulting in ectodermal dysplasia (in a random-bred cat); the CYFIP2 gene associated with urate stones (Egyptian mau); and the SH3PXD2A gene, associated with feline infectious peritonitis (random bred cat).
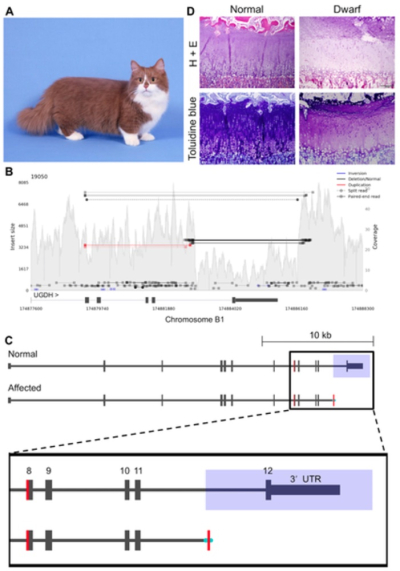
The genetic roots of dwarfish more complicated according to these researchers, involving "a complex deletion coupled with a nearby potential duplication event" on cat chromosome B1, that disrupted the gene for UDP-glucose-6-dehydrogenase (UGDH). This trait is inherited as an autosomal dominant mutant and a breed-standard for Munchkins, characterized by shortened limbs and anormal torso. Here, the scientists report that:
SV analysis within the critical region previously identified by linkage and GWAS on chromosome B1:170,786,914–175,975,857 revealed a 3.3 kb deletion at position chrB1:174,882,897–174,886,198, overlapping the final exon of UDP-glucose 6-dehydrogenase (UGDH). Upon manual inspection of this SV, a 49 bp segment from exon 8 appeared to be duplicated and inserted 3.5 kb downstream, replacing the deleted sequence. This potentially duplicated segment was flanked by a 37 bp sequence at the 5`end and a 20 bp sequence at the 3' end, both of unknown origin.
(This topography is illustrated in the Figure below.) The scientists further hypothesized that the known activity of the affected gene, UGDH, which is involved in "proteoglycan synthesis in chondrocytes" could effect this phenotype by interfering with development. Evidence for this mechanism was observed in the epiphyseal plate, which in Munchkins showed "a disorganized columnar arrangement" and proteoglycan depletion. They report that sequencing the mRNA produced by the mutant UGDH is yet to be reported, which would perhaps shed light on whether the role of this mutation was as they propose it to be.
This group published a more recent paper in March 2021, entitled "Ultracontinuous Single Haplotype Genome Assemblies for the Domestic Cat (Felis catus) and Asian Leopard Cat (Prionailurus bengalensis)," in the Journal of Heredity, 112(2): 165–73, providing additional details relating to its analysis of the domestic cat and leopard cat components of the hybrid genome.
The results reported in these papers illustrated how far we have come and perhaps how far we have to go in using genetic methods for identifying the causes and possible cures for a variety of diseases.
*Reuben M. Buckley, Brian W. Davis, Wesley A. Brashear, Fabiana H. G. Farias, Kei Kuroki, Tina Graves, LaDeana W. Hillier, Milinn Kremitzki, Gang Li, Rondo P. Middleton, Patrick Minx, Chad Tomlinson, Leslie A. Lyons, William J. Murphy, Wesley C. Warren