
Lions (Panthera leo) are a widely distributed group of terrestrial mammals, ranging during the Pleistocene (from about 2,580,000 to 11,700 years ago) in Eurasia, Africa, and North America, with species that included the still-extant modern lions (Panthera leo leo), the cave lion (Panthera leo spelaea), and the American lion (Panthera leo atrox). From the Pleistocene to modern times, all lion species except the modern lion have gone extinct (the cave and American lions in the late Pleistocene, about 14,000 years ago), and their modern range is limited to Sub-Saharan Africa and an isolated group restricted to the Kathiawar Peninsula of Gujarat State in India. In modern times, lion populations present in southwestern Eurasia were eradicated in the 19th and early 20th Centuries, and lions (particularly the Barbary lion, the Cape lion, and lion species endemic to the Middle East) disappeared through extinction from Northern Africa during the 20th century. Modern extirpation of lion populations is almost completely a consequence of human population growth, predation, and restriction of traditional environmental ranges, what is euphemistically called "anthropogenic factors."
Population studies, most relying on mitochondrial DNA (mtDNA), have been pursued for living species and remains of extinct lions, with varying degrees of confidence. Last month, an international group of researchers published a paper in the Proceedings of the National Academy of Sciences USA entitled "The evolutionary history of extinct and living lions" that reported results of genomic DNA analysis of twenty lion specimens, comprising two cave lions (Panthera leo spelaea), about 30,000 years old from Siberia and the Yukon; 12 "historic" lions (Panthera leo leo/Panthera leo melanochaita) that lived between the 15th and 20th centuries outside the current geographic distribution of lions, and 6 present-day lions from Eastern and Southern Africa (4) and India (2). The broad conclusions these authors drew from this genetic data is that the cave and modern lion species shared a common ancestor that lived about a half million years ago, and that modern lions consist of two lineages that diverged about 70,000 years ago. The "orphan" Indian lions showed what the authors termed "a nearly complete absence of genetic diversity," consistent with their low population sizes in recent years.
The whole genome studies verified the inference from earlier mtDNA studies that the cave lions were a genetic "outgroup" of modern lions. The two lineages of modern lions that diverged ~70,000 years ago comprised a "northern" lineage of lions from Asia, North Africa and West Africa, and a second "southern" lineage from Central, East and South Africa. But unlike earlier mtDNA results, the genomic analyses showed that Central African lions were closely related to southern rather than northern African lions. These authors' results also supported a closer but geographically incongruous relationship between North African lions with Asiatic lions rather than West African lions (which the authors say is "not unusual" in the cat family.
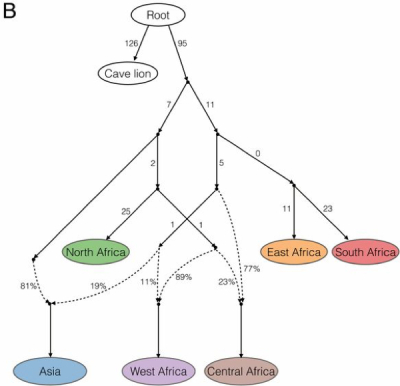
Fig. 3(B) from de Manuel et al., 2020.
Using a series of different genetic analytical techniques, these authors report a consistent time of divergence between ancestral lion populations and modern lions of about 500,000 year ago; using derived allele statistics this estimate was found to be 470,000 y-ago (392,000 – 529,000), whereas using pseudodiploid genomes (from the X chromosome of a male specimen) and "the estimate[d] rates of coalescence between their ancestral populations," this estimate was found to be 495,00 y-ago (460,000 – 578,000), and using sequence divergence an estimate of 540,000 y-ago (which relied upon a mutation rate of 4.5 × 10−9 per generation≤ a population size of 55,000 individuals, and a "transition (A-G or C-T)/transversion (A/G-C/T) ratio of 1.9," resulting in 108,000 generations at a generation time of ~5 years). The authors state, based on the different methods of estimating the divergence time, that "[g]iven the general congruence among our estimates, we conclude that the most likely split time between cave and modern lions is ca. 500,000 y ago," which they note is "remarkably consistent with the early Middle Pleistocene appearance of P. fossilis in the European fossil record."
The paper next provides a genetic analysis of gene flow (i.e., interbreeding) between cave lions and modern lions in their overlapping range in southwestern Eurasia during the Pleistocene; their results showed no evidence of gene flow between the Siberian cave lion and modern lions. In contrast, these researchers found evidence of interbreeding between Yukon cave lions and South African modern lions, which further analysis appeared to be an artifact of the differences in "depth of [genomic sequence] coverage between the Yukon and Siberian cave lions. The authors conclude that "there is no robust evidence for gene flow between the cave lion populations represented by our two samples and any of the modern lion lineages tested" (which would be expected for divergent species). The authors speculate that these results could be due to a lack of sympatry between cave and modern lions, of sexual selection due to the cave lion lacking the characteristic prominent mane exhibited by modern lions.
The authors also report the results of principal component analyses on the modern lion specimens, which were consistent with the "distinctiveness" of the southern and northern African lion populations. Repeating some of the comparative analyses performed between cave and modern lions, the authors report a divergence between these African lions of ~70,000 years (52,000-98,000 y-ago), which the authors consider to be consistent with earlier mtDNA-derived estimates and also reflect a "severe" population bottleneck in the northern Africa lineage occurring coincident with a sharp reduction in the population of this lineage at about the same time. (Both cheetahs and humans have experienced such population bottlenecks in their evolutionary histories.) These authors report that there is also evidence for genetic admixture between populations of African lions, particularly in Central African lion populations. They describe the Central African lion population as a potential "melting pot" of lineages after their divergence ~70,000 year ago. Another anomalous result, evidence of close genetic relationships between southern African lions ad Asiatic lions (e.g., with about 18.5% of Asiatic lions genetic ancestry arising from southern African lion populations), these authors postulate might have arisen from "migration corridors between Sub-Saharan African and the Near East may have existed in the past, for instance through the Nile basin in the early Holocene" and that such interbreeding with northern African lions and Asiatic lions having been prevented by "geographical barriers represented by the Atlas Mountains and the Sahara desert."
Among modern lions, assessment of autosomal heterozygosity frequency and the prevalence of "runs of homozygosity" were "consistent with a population history of consecutive bottlenecks in the northern lineage as their ancestors migrated away from Sub-Saharan Africa and persisted in more isolated smaller populations," which the authors speculate might also have arisen from "sustained anthropogenic pressure" due to a possible correlation between these effects on lion populations of large human populations in the Indus Valley and Mesopotamia in Asia, and Egyptian, Greek and Roman empires in North Africa. The capacity of these literally man-made effects on lion populations are also seen in a reduction in genetic diversity in southern African lion populations associated with European colonization during the 20th Century.
Turning to the Indian lions, the authors report the greatest extent of population reduction and inbreeding in these lions (a 16-fold reduction in heterozygosity compared with modern southern African lions), with 90% of the Indian lion genomic DNA residing in ROHs. Further analyses showed that Indian lions carry on average 12/7% more deleterious mutations in homozygosity, which results in a substantial genetic load and even more so should these mutations be recessive (i.e., individuals would carry both mutant copies of the gene).
The authors conclude their paper with a section on "implications for conservation," including efforts to resuscitate extinct or nearly extinct populations a la Jurassic Park. (The risks of this embodiment of "playing God" using genetic methodologies have been set forth in Beth Shapiro's book, How to Clone a Mammoth: The Science of De-extinction.) These authors caution that "although conservation efforts are contributing to increasing population size after centuries of decline, their remarkable lack of genomic diversity suggests that they could be extremely susceptible to inbreeding depression and genetic erosion, as well as future pathogen outbreaks."
While providing the first whole-genome sequencing comparison of several extinct and living lion species, the paper illustrates that significant additional work will be needed to sort out the interrelationships between these difference forms of the King of Beasts.
* Marc de Manuel, Ross Barnett, Marcela Sandoval-Velasco, Nobuyuki Yamaguchi, Filipe Garrett Vieira, M. Lisandra Zepeda Mendoza Shiping Liu, Michael D. Martin, Mikkel-Holger S. Sinding, Sarah S. T. Mak, Christian Carøe, Shanlin Liu, Chunxue Guo, Jiao Zheng, Grant Zazula, Gennady Baryshnikov, Eduardo Eizirik, Klaus-Peter Koepfli, Warren E. Johnson, Agostinho Antunes, Thomas Sicheritz-Ponten, Shyam Gopalakrishnan, Greger Larson, Huanming Yang, Stephen J. O'Brien, Anders J. Hansen, Guojie Zhang, Tomas Marques-Bonet, and M. Thomas P. Gilbert
Institutions: PRBB, Barcelona, Spain; University of Copenhagen; University Malaysia Terengganu, 21030 Kuala Nerus, Terengganu, Malaysia; University of Birmingham; eBGI-Shenzhen, China; Norwegian University of Science and Technology; University of Chinese Academy of Sciences; Yukon Palaeontology Program Russian Academy of Sciences; Pontifícia Universidade Católica do Rio Grande do Sul (PUCRS), Brazil; INCT-EECBio), Brazil; Instituto Pró-Carnívoros, Brazil; Smithsonian Institution; Walter Reed Army Institute of Research; University of Porto, Portugal; qDepartment of Biology, Faculty of Sciences, University of Porto, 4169-007 Porto, Portugal; Asian Institute of Medicine, Science and Technology, Malaysia; University of Oxford, OX; James D. Watson Institute of Genome Science, Hangzhou, China; Information Technologies, Mechanics and Optics University, Wt. Petersburg, Russia; Nova Southeastern University, Ft. Lauderdale, FL;, University of Copenhagen, 1350 Copenhagen, Denmark; Kunming Institute of Zoology, Chinese Academy of Sciences,Kunming, China; The Barcelona Institute of Science and Technology; ICREA, Barcelona, Spain; and Universitat Autònoma de Barcelona, Barcelona, Spain
Photo Credit: Image of Lion (Panthera leo) lying down in Namibia by Kevin Pluck, from the Wikimedia Commons under the Creative Commons Attribution 2.0 Generic license.