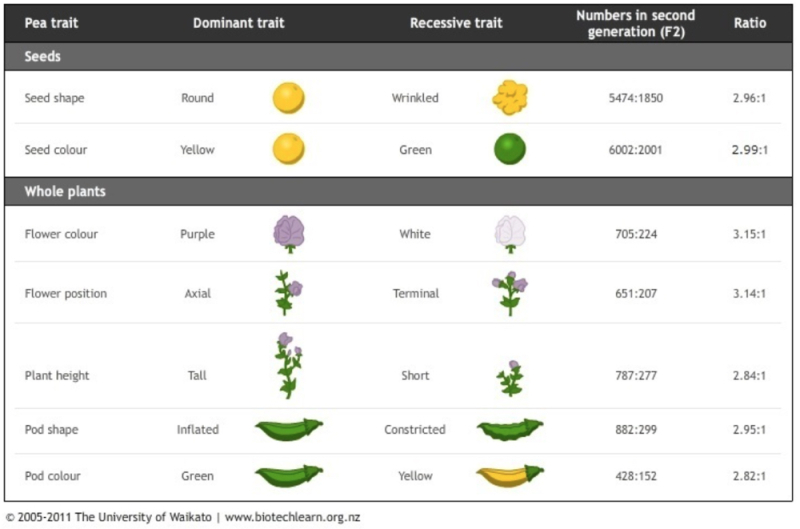
Gregor Mendel's great good fortune (or extraordinary prescience) was that he chose for the traits he used to illustrate the genetic control of inheritance (despite having no inkling of its mechanism) traits in his pea plants that were controlled by discrete genes (yellow v green, wrinkled v smooth, tall v short).
Mendelian inheritance exists in most species (ABO blood type in humans, for example), but many traits are multivariate (i.e., being caused by expression of several genes). While these include susceptibility to many diseases like cancer, the most common multivariate trait is height, which varies widely in different human ethnic and racial groups.
Recently an international group of genomics researchers* published a paper in Nature entitled "A positively selected FBN1 missense variant reduces height in Peruvian individuals." The interest in this Peruvian population is that they are recognized as being among the shortest known existing human group that is otherwise ethnically diverse (165.3 cm or 5'5" for men and 152.9 cm or 5' for women). The FBN1 gene variant reported by these researchers was found to have a specific aspartic acid residue changed to a glycine residue (E1297G, due to a change from a transition mutation of a T to a C) correlated with an average reduction in height of 2.2 cm in heterozygotes and 4.4 cm in homozygotes (corresponding to 0.87 in and 1.75 in, respectively). The FBN1 gene encodes extracellular matrix protein fibrillin 1, a major structural component of myofibrils. Individuals homozygous for the variant have less densely packed fibrillin-1-rich microfibrils with irregular edges in skin tissues.
Population genetic studies revealed that non-African populations showed the variant to be under positive selection (i.e., the variant conferred a selective advantage) in the Peruvian population studied, preferentially associated with coastal rather than mountain environments. The cohort in this study comprised 3,134 subjects / 1,947 households, that found negative statistical correlation between height and Native American ancestry.
Genome-wide association studies were performed, and five highly linked single-nucleotide polymorphisms (SNP) were found that overlapped at the chromosomal location of FBN1 (15q21.1). One of these produces the missense variant (E1297G) in the coding sequence of the FBN1 protein (fibrillin); the other SNPs are located in introns. The variant in the coding sequence was localized to the neonatal region of the protein, specifically the calcium-binding epidermal growth factor (cbEGF) domain (which have been associated with Marfan syndrome and the like, which paradoxically are characterized by greater than normal height). This is in contrast to all other height-related FBN1 mutants, which localized to the TGFβ-binding domains. Molecular analysis showed that this variant was adjacent to residues known to be involved in calcium binding. The authors conceded that "[u]nderstanding the cellular mechanisms that connect FBN1(E1297G) to microfibril structures and height requires further functional studies."
These results were also obtained when genotypes from 598 individual Peruvians were obtained. Similar results were also obtained when height information from databases (Genetic Investigation of Anthropometric Traits, GIANT) and Population Architecture using Genomics and Epidemiology, PAGE).
For comparison, previous studies on predominantly European populations (n~700,000 individuals) "have identified 3,290 independent common height-associated variants." These height-associated variants explained only about 25% (6.1%) of height variants found in European populations (24.6%). There was some evidence that these predictive differences were related to population structure and the relatively lower proportion of European ancestors in the Peruvian population.
Another difference between these European genome-identified height variants is that these show on average of difference on 0.5 cm per variant allele, while the FBN1 variants identified in the Peruvian population had on average a 2.2 cm difference in height of individual bearing these genetic variants. Statistical analyses indicated that this variant had been the subject of positive selection. Alternative explanations were not persuasive; for example, the FBN1 locus is located only 266 kilobases (kb) away from SLC24A5 gene, known to be under positive selection because of its involvement in skin pigmentation; but there was no linkage found between variants in FBN1 and SCL24A5.
FBN1 variants and their patterns of inheritance were also examined from three geographically separated Peruvian populations: coastal, Andes, and Amazon. These analyses showed that the frequencies of the variant allele in these populations was 9.7%, 1.7%, and 0%, respectively. The authors state that these differences were extreme, appearing in only 0.7% of all variants (where n = 9,381,550). Looking more closely at subpopulations, the frequency of the variant FBN1 allele is higher in the coastal Moches population, who are "far below" average height for Peruvian ("158 cm and 147 cm for Moches men and women versus 164 cm and 152 cm for Peruvian men and women measured in the same year").
The paper also addresses the molecular biology of the genetic change, noting that the changes genetic sequence (T -> C) resulted in substitution of "a large, negatively charged glutamic acid for a glycine, the smallest amino acid" in the encoded protein. This protein is "an extracellular matrix glycoprotein that serves as the structural backbone of force-bearing microfibrils in elastic and non-elastic tissues and is also involved in tissue development, homeostasis and repair by interacting with transforming growth factor (TGFβ) and other growth factors." Besides its effect on height there is no known association of this variant with any human disease. This is in contrast with other mutations in the gene, which are associated with nine "dominantly inherited Mendelian diseases" that include "skeletal anomalies and changes in skin elasticity." A physiological analysis of 11 individuals in the studied cohort ("2 homozygous (C/C) individuals, 2 heterozygous (C/T) individuals and 7 matched controls with the reference (T/T) genotype") showed no frank skeletomuscular abnormalities, while one individual (having the C/C genotype) "had a notably thicker skin as assessed in a total body skin examination and appeared older than the stated age," where another individual with this genotype did not. The authors also reported that "[s]kin biopsies, on the other hand, showed shorter microfibrillar projections from the dermal–epidermal junction into the superficial (papillary) dermis as well as less fibrillin 1 deposition in the deeper dermis." And electron microscopic analysis showed "individuals with the C/C genotype have less densely packed microfibrils with irregular edges and with microfibrils embedded in less dense collagen bundles, confirming the abnormal appearance of fibrillin 1 observed in immunohistochemical analysis of the skin biopsies."
The paper closes with an acknowledgement that understanding selection of this genetic variant in this population "is a challenging task and requires further investigation" and:
In addition to its implications in medical and population genetics, this study highlights the importance of large-scale genetic studies in underrepresented and founder populations. Our findings show that genetic studies in different populations can uncover previously undescribed trait-associated variants with large effects in functionally relevant genes. Similar studies in diverse populations are required to capture the extent of human genetic diversity and to expand the benefits of genetic research to all human populations.
* Samira Asgari, Yang Luo, Ali Akbari, Gillian M. Belbin7, Xinyi Li, Daniel N. Harris, Martin Selig, Eric Bartell, Roger Calderon, Kamil Slowikowski Carmen Contreras, Rosa Yataco, Jerome T. Galea, Judith Jimenez, Julia M. Coit, Chandel Farroñay, Rosalynn M. Nazarian12, Timothy D. O'Connor, Harry C. Dietz, Joel N. Hirschhorn, Heinner Guio, Leonid Lecca, Eimear E. Kenny, Esther E. Freeman, Megan B. Murray & Soumya Raychaudhuri from Harvard Medical School, the Broad Institute/MIT, Mt Sinai School of Medicine, University of Maryland Medical School, University of Pennsylvania, Boston Children's Hospital, Socios en Salud, Lima, Peru, University of South Florida, Johns Hopkins University School of Medicine, University of Manchester, UK, and Instituto Nacional de Salud, Lima, Peru.